suppressPackageStartupMessages({
library(EasyProtein)
library(tidyverse)
library(SummarizedExperiment)
})Quick Start with EasyProtein
1. Tutorial metadata
-
Tutorial name: Quick Start with EasyProtein
-
Tutorial type: Minimal end-to-end workflow
-
Target audience: First-time users of
EasyProtein
-
Primary goal: Show how to complete a basic
proteomics analysis workflow with minimal input and minimal parameter
tuning
-
Expected outcome: You will learn how to:
- load example data
- create a standard EasyProtein analysis object
- inspect data quality
- visualize sample structure
- perform a simple two-group comparison (Liver, 1w vs 8w, male only)
- interpret core outputs
2. Load example data and build
SummarizedExperiment
This tutorial uses the file
inst/extdata/mouse_multi-organ_DIA_report.pg_matrix.tsv.gz.
exp_file <- system.file(
"extdata",
"mouse_multi-organ_DIA_report.pg_matrix.tsv.gz",
package = "EasyProtein"
)
stopifnot(file.exists(exp_file))
se_obj <- rawdata2se(exp_file)
#> [rawdata2se] Reading input took 1.78 seconds
#> [rawdata2se] Detecting raw columns took 0.00 seconds
#> [rawdata2se] Preprocessing took 1.71 seconds
#> [rawdata2se] Missing-value filtering took 94.31 seconds
#> [rawdata2se] Imputation took 8.05 seconds
#> [rawdata2se] Building SummarizedExperiment took 2.09 seconds
#> [rawdata2se] Stability filtering took 1.14 seconds
#> [rawdata2se] Done took 0.00 seconds
se <- se_obj$se
se
#> class: SummarizedExperiment
#> dim: 11497 299
#> metadata(0):
#> assays(4): raw_intensity intensity conc zscale
#> rownames(11497): 0610012G03Rik 1700010I14Rik ... mt-Nd3 rp9
#> rowData names(1): gene
#> colnames(299): brain_1w_F_5 brain_1w_F_7 ... Muscle8w_M_4 Muscle8w_M_5
#> colData names(4): sample condition rep group3. Add sample metadata
rawdata2se() already creates basic
condition and rep. Here we derive
tissue, age, and sex from
condition as requested.
se$tissue <- str_extract(se$condition,'[A-Za-z]+')
se$age <- str_extract(se$condition,'\\dw')
se$sex <- str_extract(se$condition,'\\w$')
as.data.frame(SummarizedExperiment::colData(se)) %>%
dplyr::select(sample, condition, rep, tissue, age, sex) %>%
head()
#> sample condition rep tissue age sex
#> brain_1w_F_5 brain_1w_F_5 brain_1w_F 5 brain 1w F
#> brain_1w_F_7 brain_1w_F_7 brain_1w_F 7 brain 1w F
#> brain_1w_F_8 brain_1w_F_8 brain_1w_F 8 brain 1w F
#> brain_1w_F_10 brain_1w_F_10 brain_1w_F 10 brain 1w F
#> brain_1w_M_1 brain_1w_M_1 brain_1w_M 1 brain 1w M
#> brain_1w_M_3 brain_1w_M_3 brain_1w_M 3 brain 1w M4. Quick QC
We inspect three basic QC aspects:
- sample intensity distribution
- missing values per sample
- detected protein number per sample
keep <- se$tissue == "live" &
se$sex == "M" &
se$age %in% c("1w", "8w")
se_sub <- se[, keep]
p_density <- plotSE_density(se_sub)
p_missing <- plotSE_missing_value(se_sub)
p_detected <- plotSE_protein_number(se_sub)
p_density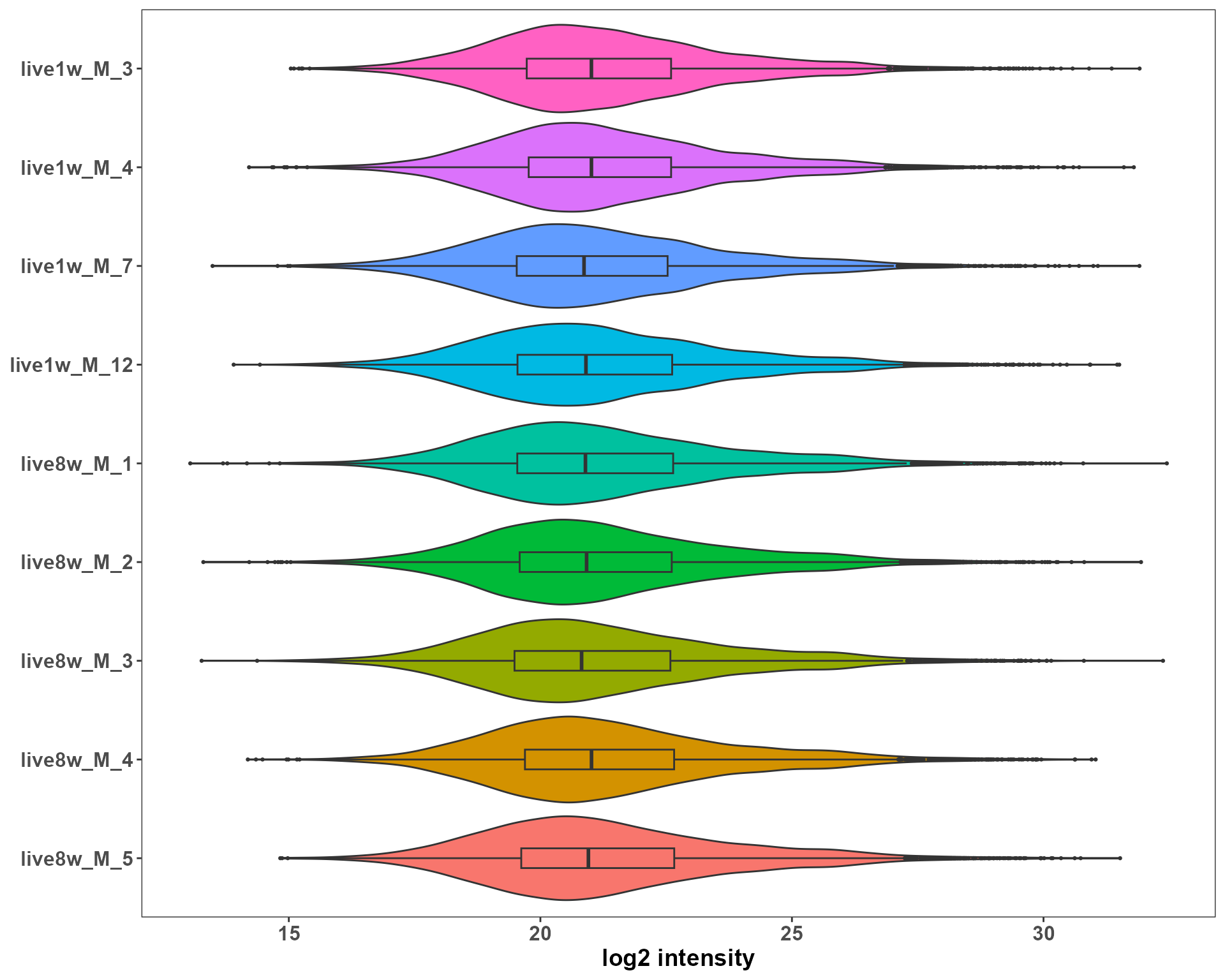
p_missing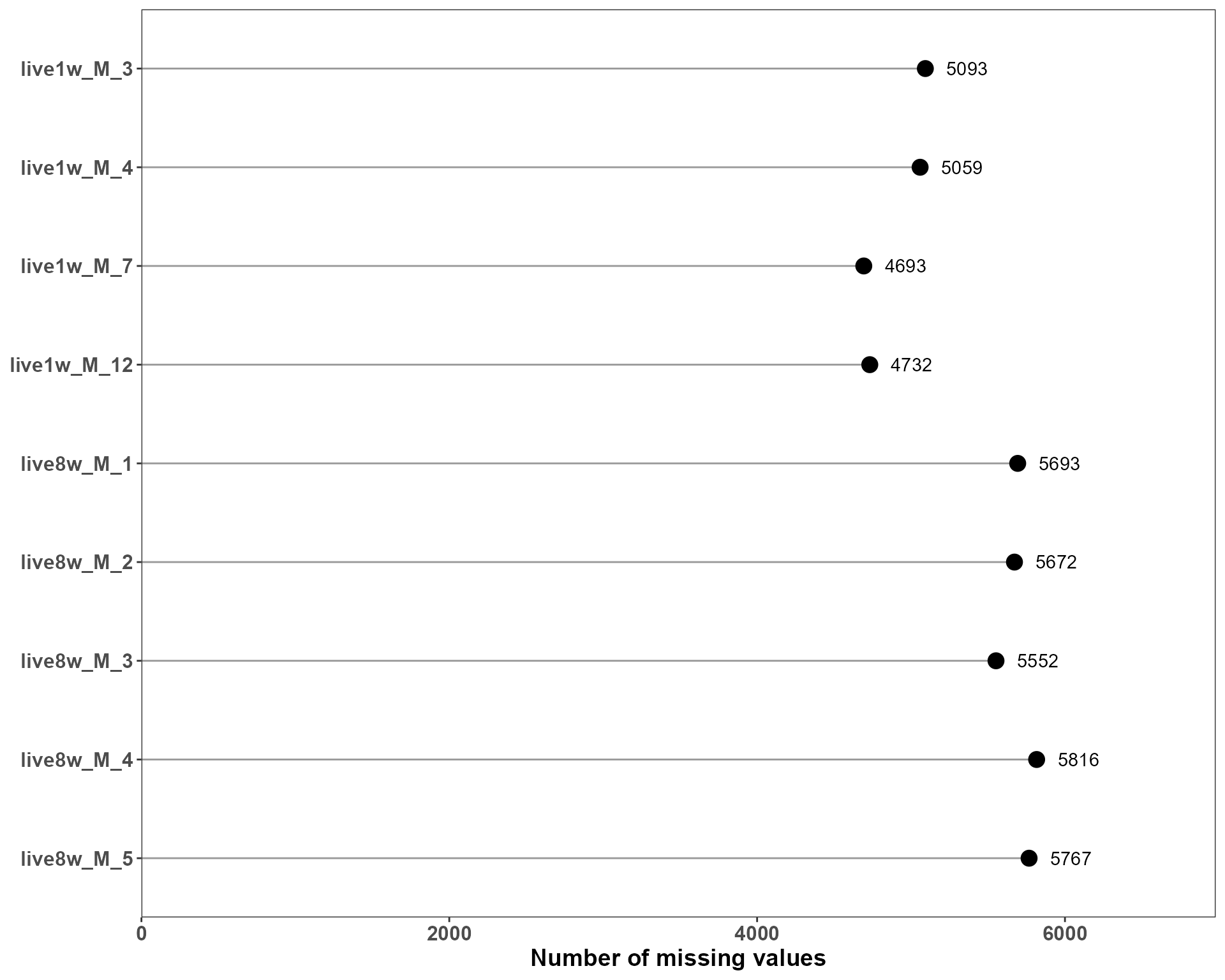
p_detected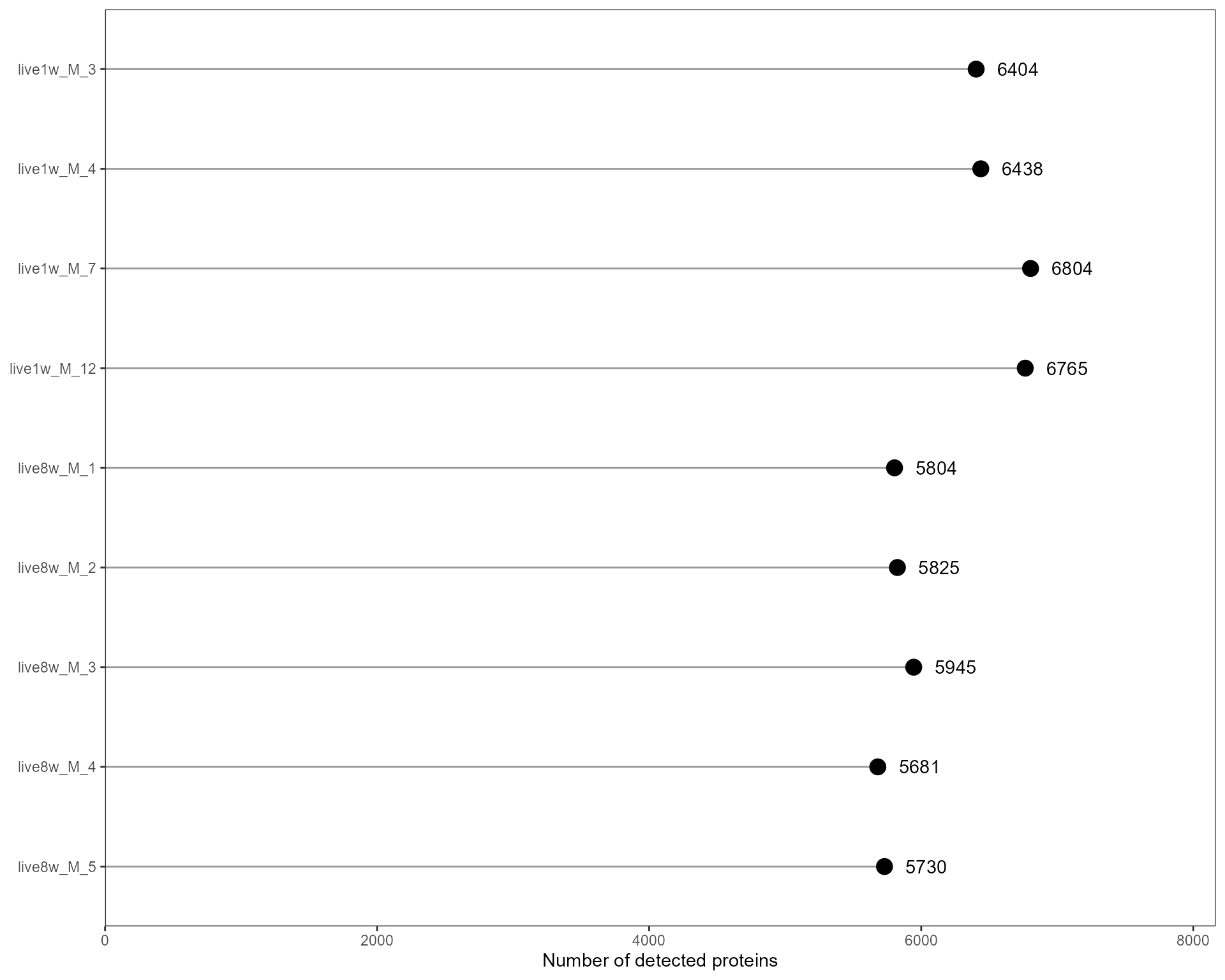
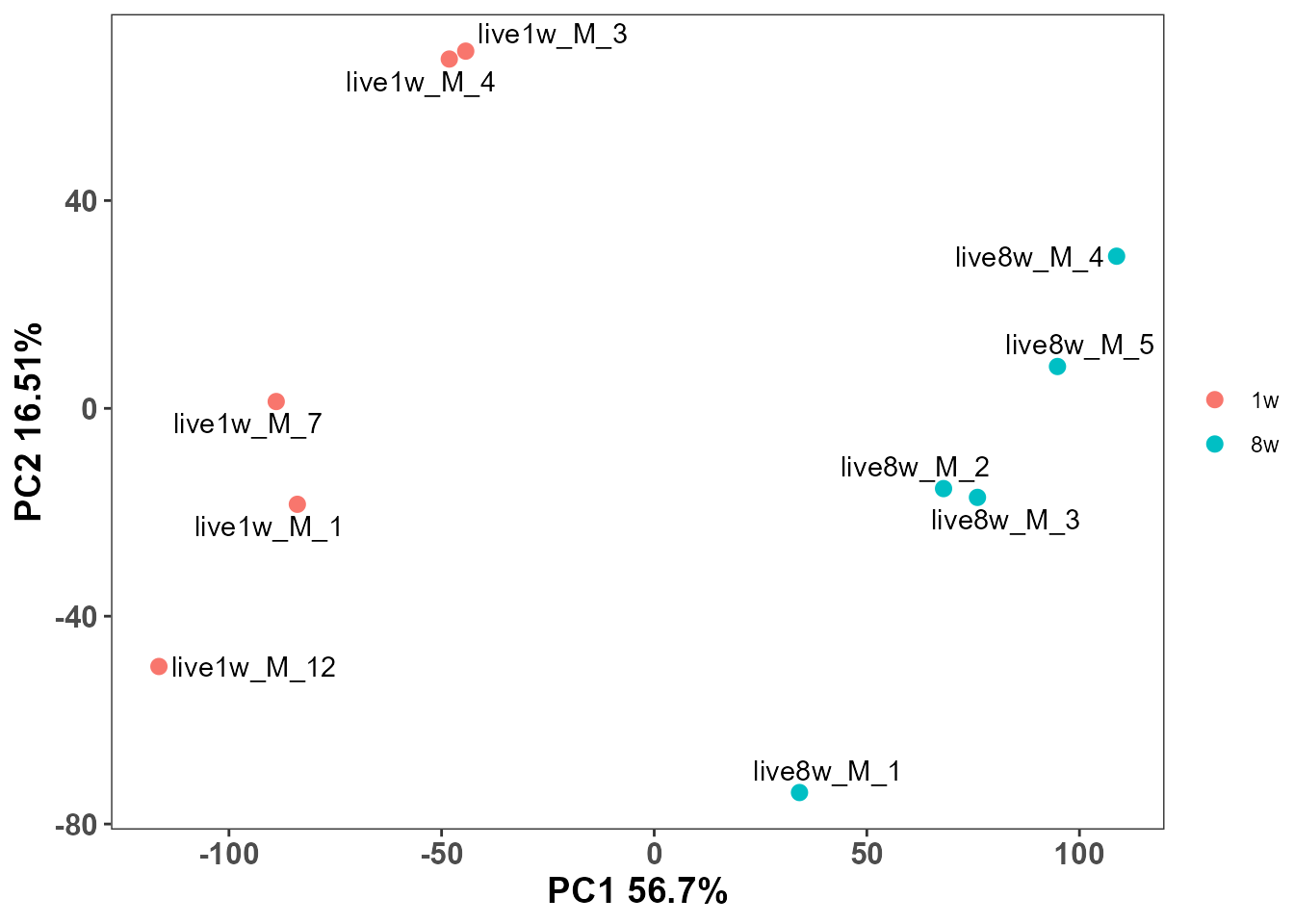
5. Visualize sample structure with PCA
Use concentration matrix (assay = "conc") and color by
tissue.
conc_mtx <- SummarizedExperiment::assay(se_sub, "conc")
pca.res <- FactoMineR::PCA(t(conc_mtx), graph = FALSE)
pca.df <- as.data.frame(pca.res$ind$coord)
pca.df$sample <- rownames(pca.df)
meta_df <- as.data.frame(SummarizedExperiment::colData(se_sub))
pca.df <- dplyr::left_join(pca.df, meta_df, by = "sample")
plot_pca(
pca.df = pca.df,
pca.res = pca.res,
colorby = "age",
label = "sample"
)
6. Two-group comparison: Liver, 1w vs 8w, male only
We subset samples to:
tissue == "Liver"sex == "m"age %in% c("1w", "8w")
Then run unpaired limma-based DEG analysis via
se2DEGs().
deg_res <- se2DEGs(
se = se_sub,
compare_col = "age",
ref = "1w",
cmp = "8w",
logFC_cutoff = 1,
adj_p_cutoff = 0.05
)
head(deg_res)
#> gene live1w_M_1 live1w_M_3 live1w_M_4 live1w_M_7 live1w_M_12
#> 1 0610012G03Rik 3.4170122 2.7307321 3.5057499 4.4052290 3.345071
#> 2 1700010I14Rik 0.9392232 0.9654005 0.9636142 0.9366667 0.919449
#> 3 1810009J06Rik 0.4087430 3551.7545240 1793.9272352 38.1159646 3649.433156
#> 4 1810024B03Rik 0.9392232 0.9654005 0.9636142 0.9366667 0.919449
#> 5 1810065E05Rik 41.1277752 42.2740584 42.1958388 41.0158293 40.261882
#> 6 2010106E10Rik 0.9392232 0.9654005 0.9636142 0.9366667 0.919449
#> live8w_M_1 live8w_M_2 live8w_M_3 live8w_M_4 live8w_M_5 mean_log2_1w
#> 1 1.8695419 1.9201328 1.929792 1.9712530 1.9541805 1.80002963
#> 2 0.9446468 0.9702095 0.975090 0.9960396 0.9874132 -0.02024861
#> 3 3016.5522357 3098.1819292 3113.766947 3180.6656198 3153.1188707 7.70778304
#> 4 0.9446468 0.9702095 0.975090 0.9960396 0.9874132 -0.02024861
#> 5 0.9446468 0.9702095 0.975090 0.9960396 0.9874132 5.37188591
#> 6 0.9446468 0.9702095 0.975090 0.9960396 0.9874132 -0.02024861
#> mean_log2_8w logFC P.Value adj.P.Val median_1w median_8w
#> 1 0.97820863 -0.82182099 4.856343e-05 2.118110e-04 3.4170122 1.929792
#> 2 0.02272051 0.04296912 3.520132e-02 4.948155e-02 0.9392232 0.975090
#> 3 11.60362179 3.89583875 1.374975e-01 1.724268e-01 1793.9272352 3113.766947
#> 4 0.02272051 0.04296912 3.520132e-02 4.948155e-02 0.9392232 0.975090
#> 5 0.02272051 -5.34916541 5.591284e-15 5.541638e-13 41.1277752 0.975090
#> 6 0.02272051 0.04296912 3.520132e-02 4.948155e-02 0.9392232 0.975090
#> DEGs_types
#> 1 NS
#> 2 NS
#> 3 NS
#> 4 NS
#> 5 DOWN
#> 6 NS
table(deg_res$DEGs_types)
#>
#> DOWN NS UP
#> 1725 8983 7897. Interpret core outputs
The DEG result table contains:
-
effect size:
logFC(positive means higher in8wvs1w) -
statistical significance:
P.Value,adj.P.Val -
classification:
DEGs_types(UP,DOWN,NS) -
group summaries:
median_1w,median_8w
Minimal interpretation workflow:
- prioritize proteins with
adj.P.Val <= 0.05 - use
|logFC| >= 1to focus on stronger changes - read
median_1wandmedian_8wtogether withlogFC
top_hits <- deg_res %>%
dplyr::arrange(adj.P.Val, dplyr::desc(abs(logFC))) %>%
dplyr::select(gene, logFC, adj.P.Val, DEGs_types, starts_with("median_"))
head(top_hits, 20)
#> gene logFC adj.P.Val DEGs_types median_1w median_8w
#> 1 Csn1s1 10.182433 8.013961e-16 UP 0.9392232 1146.46359
#> 2 Cdca3 10.040292 8.013961e-16 UP 0.9392232 1038.89116
#> 3 Naaladl1 9.982767 8.013961e-16 UP 0.9392232 998.27682
#> 4 Stx1b 9.618075 9.323407e-16 UP 0.9392232 775.29011
#> 5 Mybpc1 -9.576063 9.323407e-16 DOWN 770.8484387 0.97509
#> 6 Cck 9.184286 1.558556e-15 UP 0.9392232 573.94779
#> 7 Defa21 -8.684485 3.045286e-15 DOWN 415.4870265 0.97509
#> 8 Eml6 8.494027 5.434018e-15 UP 0.9392232 353.90296
#> 9 Metrnl -8.253243 5.434018e-15 DOWN 308.1239990 0.97509
#> 10 Sytl1 8.107017 5.973509e-15 UP 0.9392232 271.98647
#> 11 Myom1 8.104425 5.973509e-15 UP 0.9392232 271.49834
#> 12 Nefh -8.101866 5.973509e-15 DOWN 277.4279338 0.97509
#> 13 Atp1b4 7.939756 7.849712e-15 UP 0.9392232 242.20809
#> 14 Eef1a2 -7.715343 1.091110e-14 DOWN 212.2146391 0.97509
#> 15 Nefl 7.687151 1.189352e-14 UP 0.9392232 203.29790
#> 16 Krt85 7.607042 1.190601e-14 UP 0.9392232 192.31482
#> 17 Pmfbp1 7.587927 1.190601e-14 UP 0.9392232 189.78299
#> 18 Krt17 7.560404 1.190601e-14 UP 0.9392232 186.19589
#> 19 Trim54 7.531913 1.190601e-14 UP 0.9392232 182.55411
#> 20 Has1 -7.481887 1.190601e-14 DOWN 180.5026100 0.975098. GO and KEGG enrichment (table output only)
Here we use significant DE proteins
(adj.P.Val <= 0.05) for over-representation analysis. We
only return result tables (no plotting).
deg_sig_genes <- deg_res %>%
dplyr::filter(adj.P.Val <= 0.05, DEGs_types %in% c("UP", "DOWN")) %>%
dplyr::pull(gene) %>%
unique()
length(deg_sig_genes)
#> [1] 2514
if (!requireNamespace("clusterProfiler", quietly = TRUE) ||
!requireNamespace("org.Mm.eg.db", quietly = TRUE)) {
message("Please install 'clusterProfiler' and 'org.Mm.eg.db' to run enrichment.")
} else {
go_res <- enrichment_analysis(
gene = deg_sig_genes,
db = "GO",
species = "mouse",
keyType = "SYMBOL",
qvalue_cutoff = 0.05
)
kegg_res <- enrichment_analysis(
gene = deg_sig_genes,
db = "KEGG",
species = "mouse",
keyType = "SYMBOL",
qvalue_cutoff = 0.05,
use_internal_data = TRUE #Select false will using online data
)
}
#>
#>
#>
#> KEGG.db contains mappings based on older data because the original
#> resource was removed from the the public domain before the most
#> recent update was produced. This package should now be considered
#> deprecated and future versions of Bioconductor may not have it
#> available. Users who want more current data are encouraged to look
#> at the KEGGREST or reactome.db packages
#> 'select()' returned 1:1 mapping between keys and columns
#> Warning in clusterProfiler::bitr(gene, fromType = keyType, toType = "ENTREZID",
#> : 2.9% of input gene IDs are fail to map...
# tables only
head(go_res, 20)
#> ONTOLOGY ID Description
#> GO:0009410 BP GO:0009410 response to xenobiotic stimulus
#> GO:0044282 BP GO:0044282 small molecule catabolic process
#> GO:0046394 BP GO:0046394 carboxylic acid biosynthetic process
#> GO:0016053 BP GO:0016053 organic acid biosynthetic process
#> GO:0006631 BP GO:0006631 fatty acid metabolic process
#> GO:1901605 BP GO:1901605 alpha-amino acid metabolic process
#> GO:0006575 BP GO:0006575 cellular modified amino acid metabolic process
#> GO:0006790 BP GO:0006790 sulfur compound metabolic process
#> GO:0006520 BP GO:0006520 amino acid metabolic process
#> GO:0170033 BP GO:0170033 L-amino acid metabolic process
#> GO:0007015 BP GO:0007015 actin filament organization
#> GO:0016054 BP GO:0016054 organic acid catabolic process
#> GO:0046395 BP GO:0046395 carboxylic acid catabolic process
#> GO:0031589 BP GO:0031589 cell-substrate adhesion
#> GO:0170041 BP GO:0170041 non-proteinogenic amino acid metabolic process
#> GO:0071466 BP GO:0071466 cellular response to xenobiotic stimulus
#> GO:0120254 BP GO:0120254 olefinic compound metabolic process
#> GO:0072330 BP GO:0072330 monocarboxylic acid biosynthetic process
#> GO:0006805 BP GO:0006805 xenobiotic metabolic process
#> GO:0006690 BP GO:0006690 icosanoid metabolic process
#> GeneRatio BgRatio RichFactor FoldEnrichment zScore pvalue
#> GO:0009410 97/2363 350/28905 0.2771429 3.390103 13.42313 1.496196e-27
#> GO:0044282 97/2363 353/28905 0.2747875 3.361292 13.31873 3.081397e-27
#> GO:0046394 90/2363 325/28905 0.2769231 3.387415 12.91465 1.257081e-25
#> GO:0016053 90/2363 328/28905 0.2743902 3.356433 12.80642 2.584532e-25
#> GO:0006631 109/2363 465/28905 0.2344086 2.867364 12.11252 3.300667e-24
#> GO:1901605 66/2363 209/28905 0.3157895 3.862842 12.39378 1.445603e-22
#> GO:0006575 64/2363 199/28905 0.3216080 3.934016 12.39215 2.100890e-22
#> GO:0006790 84/2363 321/28905 0.2616822 3.200984 11.83183 3.125835e-22
#> GO:0006520 74/2363 282/28905 0.2624113 3.209903 11.12715 8.004668e-20
#> GO:0170033 52/2363 156/28905 0.3333333 4.077444 11.49966 3.028989e-19
#> GO:0007015 101/2363 477/28905 0.2117400 2.590074 10.44835 4.083097e-19
#> GO:0016054 64/2363 241/28905 0.2655602 3.248420 10.45831 1.289033e-17
#> GO:0046395 64/2363 241/28905 0.2655602 3.248420 10.45831 1.289033e-17
#> GO:0031589 83/2363 369/28905 0.2249322 2.751446 10.10316 1.385852e-17
#> GO:0170041 34/2363 78/28905 0.4358974 5.332042 11.43100 3.394664e-17
#> GO:0071466 55/2363 191/28905 0.2879581 3.522399 10.43585 5.012711e-17
#> GO:0120254 51/2363 172/28905 0.2965116 3.627029 10.31057 1.779394e-16
#> GO:0072330 60/2363 228/28905 0.2631579 3.219035 10.03711 2.102301e-16
#> GO:0006805 43/2363 129/28905 0.3333333 4.077444 10.45235 3.642931e-16
#> GO:0006690 46/2363 149/28905 0.3087248 3.776425 10.13815 9.644999e-16
#> p.adjust qvalue
#> GO:0009410 9.069941e-24 5.731219e-24
#> GO:0044282 9.339716e-24 5.901687e-24
#> GO:0046394 2.540141e-22 1.605094e-22
#> GO:0016053 3.916858e-22 2.475029e-22
#> GO:0006631 4.001728e-21 2.528658e-21
#> GO:1901605 1.460541e-19 9.229034e-20
#> GO:0006575 1.819371e-19 1.149645e-19
#> GO:0006790 2.368602e-19 1.496699e-19
#> GO:0006520 5.391589e-17 3.406899e-17
#> GO:0170033 1.836173e-16 1.160262e-16
#> GO:0007015 2.250158e-16 1.421855e-16
#> GO:0016054 6.000738e-15 3.791816e-15
#> GO:0046395 6.000738e-15 3.791816e-15
#> GO:0031589 6.000738e-15 3.791816e-15
#> GO:0170041 1.371897e-14 8.668899e-15
#> GO:0071466 1.899191e-14 1.200083e-14
#> GO:0120254 6.345110e-14 4.009421e-14
#> GO:0072330 7.080084e-14 4.473845e-14
#> GO:0006805 1.162287e-13 7.344391e-14
#> GO:0006690 2.913267e-13 1.840869e-13
#> geneID
#> GO:0009410 Aacs/Abat/Abcc1/Adam17/Add3/Ahr/Aim2/Akr1c12/Akr1c6/Aldh1a1/Aldh3a1/Aox1/Cers1/Comt/Crot/Cybb/Cyp1a2/Cyp1b1/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp2f2/Cyp2g1/Cyp2j11/Cyp2s1/Cyp3a11/Cyp3a41b/Cyp3a44/Cyp46a1/Dnmt1/Dpep1/Fmo5/Fzd1/Gata4/Ggh/Gpx1/Gsta2/Gsta3/Gstm1/Gstm2/Gstm3/Gstm4/Gstp1/Gstp3/Hdac4/Igf2/Kcnq2/Lck/Lcn2/Lct/Mcm7/Mgst1/Mmp2/Ncam1/Nckap1l/Nos1/Nqo1/Pam/Pcna/Pde3a/Pemt/Pnp2/Prkaa2/Prkcb/Ptprm/Rap2a/Recql5/Ren1/Slc1a2/Slc1a3/Slc22a12/Smtnl1/Snca/Sod2/Sult1b1/Sult2a1/Sult2a5/Sult2a8/Tcf3/Tfrc/Tigar/Ugt2b1/Ugt2b37/Umod/Vav3/Vkorc1
#> GO:0044282 Abat/Abcd2/Abhd2/Acacb/Acadl/Acadsb/Acat1/Acat3/Acmsd/Acox3/Adh1/Adh4/Adtrp/Agxt/Ahcy/Akr1c18/Akr1c6/Aldh1a1/Aldh1a7/Aldh1l1/Aldh1l2/Aldh6a1/Arg1/Arg2/Arsb/Bcat1/Bcat2/Bdh2/Cda/Ces1f/Cpt1a/Crat/Crot/Cryl1/Cyp27a1/Cyp39a1/Cyp46a1/Cyp4f14/Dao/Dhdh/Dpep1/Echdc1/Gale/Galk1/Gcdh/Gck/Gda/Glo1/Gls2/Hal/Hao1/Hao2/Hexa/Hexb/Hgd/Hk1/Hpd/Ido2/Khk/Kmo/Ldha/Ldhd/Mgat1/Miox/Nos1/Npl/Nqo2/Nudt7/Oat/Otc/Oxct1/Pah/Pck2/Pdxp/Pfkl/Pgam2/Pkm/Pnp2/Ppat/Prodh/Prodh2/Qdpr/Renbp/Sardh/Sds/Slc16a3/Slc25a44/Sult1b1/Sult2a1/Sult2a5/Sult2a8/Tat/Tdo2/Tigar/Upb1/Upp2/Xdh
#> GO:0046394 Abat/Abcd2/Abhd2/Acacb/Acadl/Acly/Acmsd/Acsbg1/Acsm2/Acsm5/Acss1/Acss2/Agt/Agxt/Akr1c18/Akr1c6/Aldh18a1/Aldh1a1/Aldh1a3/Alox12/Alox12b/Alox12e/Alox5ap/Anxa1/Apoa4/Apoc3/Asl/Asns/Ass1/Baat/Bcat1/Bcat2/Bhmt2/Ces1f/Chst14/Cth/Cyp27a1/Cyp39a1/Cyp4a12a/Cyp7b1/Degs1/Elovl2/Fabp5/Fads1/Fads2/Fasn/Gamt/Gatm/Ggt1/Gls2/Glul/Gstm4/Gstp1/Gstp3/Haao/Hacd2/Hao1/Kmo/Ldha/Ldhb/Lpgat1/Lpl/Ltc4s/Mthfd1l/Mthfr/Myo5a/Nags/Otc/Pah/Pdk4/Phgdh/Pkm/Plod2/Prkaa1/Prkaa2/Prmt3/Prox1/Prxl2b/Psat1/Psph/Pycr2/Qki/Rbp1/Rdh9/Scd1/Sds/Slc1a3/Srr/Ugdh/Upb1
#> GO:0016053 Abat/Abcd2/Abhd2/Acacb/Acadl/Acly/Acmsd/Acsbg1/Acsm2/Acsm5/Acss1/Acss2/Agt/Agxt/Akr1c18/Akr1c6/Aldh18a1/Aldh1a1/Aldh1a3/Alox12/Alox12b/Alox12e/Alox5ap/Anxa1/Apoa4/Apoc3/Asl/Asns/Ass1/Baat/Bcat1/Bcat2/Bhmt2/Ces1f/Chst14/Cth/Cyp27a1/Cyp39a1/Cyp4a12a/Cyp7b1/Degs1/Elovl2/Fabp5/Fads1/Fads2/Fasn/Gamt/Gatm/Ggt1/Gls2/Glul/Gstm4/Gstp1/Gstp3/Haao/Hacd2/Hao1/Kmo/Ldha/Ldhb/Lpgat1/Lpl/Ltc4s/Mthfd1l/Mthfr/Myo5a/Nags/Otc/Pah/Pdk4/Phgdh/Pkm/Plod2/Prkaa1/Prkaa2/Prmt3/Prox1/Prxl2b/Psat1/Psph/Pycr2/Qki/Rbp1/Rdh9/Scd1/Sds/Slc1a3/Srr/Ugdh/Upb1
#> GO:0006631 Aacs/Abcd2/Abhd2/Acacb/Acadl/Acadsb/Acat1/Acat3/Acbd5/Acly/Acnat1/Acnat2/Acot3/Acox3/Acsbg1/Acsf2/Acsm2/Acsm5/Acss1/Acss2/Adh4/Adtrp/Agt/Ahr/Akr1c18/Akr1c6/Aldh1l2/Alox12/Alox12b/Alox12e/Alox5ap/Anxa1/Apoa4/Apoc3/Atp6v1b1/Baat/Bdh2/C3/Ces1f/Ces2b/Ces2c/Ces2g/Comt/Cpt1a/Cpt1b/Crat/Crot/Cryl1/Cyp1a2/Cyp1b1/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp2f2/Cyp2g1/Cyp2j11/Cyp2s1/Cyp4a12a/Cyp4f14/Degs1/Echdc1/Elovl2/Fabp3/Fabp4/Fabp5/Fads1/Fads2/Fasn/Gcdh/Gk/Gpx1/Gstm4/Gstp1/Gstp3/Hacd2/Hao1/Hao2/Hpgd/Lpgat1/Lpl/Ltc4s/Mblac2/Myo5a/Nudt7/Pam/Pck2/Pdk3/Pdk4/Ppargc1a/Prkaa1/Prkaa2/Prxl2b/Ptgr1/Qki/Scd1/Slc27a3/Snca/Them5
#> GO:1901605 Acadsb/Acat1/Acmsd/Agxt/Ahcy/Aldh18a1/Aldh6a1/Arg1/Arg2/Asl/Asns/Aspg/Ass1/Baat/Bcat1/Bcat2/Bhmt2/Comt/Cps1/Cth/Ctps1/Ctps2/Dao/Dglucy/Dpep1/Ero1a/Gamt/Gfpt1/Ggt1/Gls2/Glul/Glyat/Gnmt/Gpt2/Haao/Hgd/Hpd/Icmt/Ido2/Kmo/Kyat1/Kyat3/Mthfr/Nags/Nos1/Oat/Otc/P4ha1/Pah/Pemt/Phgdh/Plod2/Ppat/Prdx4/Prodh/Prodh2/Psat1/Psph/Pycr2/Qdpr/Sardh/Sds/Slc7a7/Srr/Tat/Tdo2
#> GO:0006575 Acadl/Ahcy/Aldh1l1/Aldh1l2/Asl/Ass1/Bhmt2/Ckb/Ckm/Ckmt1/Cpt1a/Cpt1b/Crat/Crot/Cth/Ctsk/Dpep1/Ero1a/Ethe1/Folh1/Gamt/Gatm/Gch1/Ggh/Ggt1/Glo1/Gnmt/Gpx1/Gsta2/Gsta3/Gstk1/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Icmt/Idh1/Lpcat4/Mgst1/Mthfd1l/Mthfr/Oplah/P4ha1/Pcyox1l/Pemt/Plod2/Prdx4/Reln/Sardh/Slc16a10/Slc1a2/Slc22a5/Sod2/Sult1b1/Sult2a1/Sult2a5/Sult2a8/Tmlhe/Tyms/Vnn3
#> GO:0006790 Acacb/Acadsb/Acat1/Acly/Acnat1/Acnat2/Acot3/Acp3/Acsm2/Acsm5/Acss1/Acss2/Agxt/Ahcy/Arsb/B3gat3/Baat/Bhmt2/Chst14/Comt/Cps1/Cs/Cth/Dnmt1/Dpep1/Ethe1/Fasn/Gal3st2/Gamt/Gcdh/Ggt1/Glce/Glo1/Glyat/Gnmt/Gpc1/Gpx1/Gsta2/Gsta3/Gstk1/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Hexa/Hexb/Hlcs/Hmgcs1/Hmgcs2/Hnmt/Hyal1/Icmt/Idh1/Mgst1/Mpc1/Mthfr/Nudt7/Oplah/Papss1/Pdk3/Pdk4/Pemt/Phgdh/Slc1a2/Slc27a3/Snca/Sod2/Stat5a/Sulf1/Sult1b1/Sult1c2/Sult2a1/Sult2a5/Sult2a8/Suox/Tdo2/Them5/Tpst1/Ugdh/Vangl2
#> GO:0006520 Abat/Acadsb/Acat1/Acmsd/Agxt/Ahcy/Aldh18a1/Aldh1a1/Aldh6a1/Arg1/Arg2/Asl/Asns/Aspg/Ass1/Baat/Bcat1/Bcat2/Bhmt2/Comt/Cps1/Cth/Ctps1/Ctps2/Dao/Ddc/Dglucy/Dpep1/Ero1a/Gamt/Gfpt1/Ggt1/Gls2/Glul/Glyat/Gnmt/Gpt2/Haao/Hgd/Hpd/Icmt/Ido2/Kmo/Kyat1/Kyat3/Mthfr/Nags/Nos1/Oat/Otc/P4ha1/Pah/Pemt/Phgdh/Plod2/Pm20d1/Ppat/Prdx4/Prodh/Prodh2/Psat1/Psph/Pycr2/Qdpr/Sardh/Sds/Slc1a3/Slc25a44/Slc7a7/Srr/Tat/Tdo2/Upb1/Wars2
#> GO:0170033 Agxt/Ahcy/Aldh18a1/Arg1/Arg2/Asl/Asns/Aspg/Ass1/Baat/Bhmt2/Cps1/Cth/Ctps1/Ctps2/Dao/Dglucy/Ero1a/Gamt/Gfpt1/Ggt1/Gls2/Glul/Glyat/Gnmt/Gpt2/Hgd/Hpd/Icmt/Kmo/Kyat1/Mthfr/Nags/Nos1/Oat/Otc/P4ha1/Pah/Pemt/Phgdh/Plod2/Ppat/Prdx4/Prodh/Prodh2/Psat1/Psph/Pycr2/Qdpr/Sds/Srr/Tat
#> GO:0007015 Abl2/Add2/Add3/Aif1/Alms1/Apoa1/Arhgap18/Arhgef15/Arhgef2/C9orf72/Cap2/Capg/Ccdc88a/Cit/Clasp1/Coro1a/Cotl1/Cyrib/Dbn1/Diaph3/Dmtn/Dpysl3/Elmo1/Elmo2/Eln/Evl/Ezr/Fchsd2/Fhod1/Flna/Fscn1/Gmfg/Gsn/Hcls1/Hdac6/Hip1r/Iqgap1/Itgb5/Jmy/Kirrel1/Limch1/Limd2/Limk1/Marcks/Marcksl1/Myadm/Myh10/Myo1a/Myo5a/Myo5b/Myo5c/Myo7b/Naa80/Nck2/Nckap1l/Nebl/Nf2/Pacsin1/Pak1/Pak3/Pdlim4/Pdxp/Pfn2/Pfn3/Pls1/Pls3/Pof1b/Ppp1r9a/Ppp1r9b/Prex1/Prkci/Prox1/Rac2/Rhoc/Rhog/Rhoj/Rhpn2/Rictor/Scin/Serpinf2/Sh3kbp1/Shroom3/Sirpa/Smad3/Specc1l/Spta1/Sptb/Src/Ssh2/Stmn1/Synpo/Synpo2l/Tagln2/Tmod2/Tmsb4x/Tpm4/Trim27/Vasp/Vil1/Zeb2/Zyx
#> GO:0016054 Abat/Abcd2/Abhd2/Acacb/Acadl/Acadsb/Acat1/Acat3/Acmsd/Acox3/Adtrp/Agxt/Ahcy/Aldh1l1/Aldh1l2/Aldh6a1/Arg1/Arg2/Arsb/Bcat1/Bcat2/Bdh2/Ces1f/Cpt1a/Crat/Crot/Cryl1/Cyp39a1/Cyp4f14/Dao/Dpep1/Echdc1/Gcdh/Gls2/Hal/Hao1/Hao2/Hexa/Hexb/Hgd/Hpd/Ido2/Kmo/Ldha/Ldhd/Nos1/Npl/Nudt7/Oat/Otc/Pah/Pck2/Ppat/Prodh/Prodh2/Qdpr/Renbp/Sardh/Sds/Slc16a3/Slc25a44/Sult2a8/Tat/Tdo2
#> GO:0046395 Abat/Abcd2/Abhd2/Acacb/Acadl/Acadsb/Acat1/Acat3/Acmsd/Acox3/Adtrp/Agxt/Ahcy/Aldh1l1/Aldh1l2/Aldh6a1/Arg1/Arg2/Arsb/Bcat1/Bcat2/Bdh2/Ces1f/Cpt1a/Crat/Crot/Cryl1/Cyp39a1/Cyp4f14/Dao/Dpep1/Echdc1/Gcdh/Gls2/Hal/Hao1/Hao2/Hexa/Hexb/Hgd/Hpd/Ido2/Kmo/Ldha/Ldhd/Nos1/Npl/Nudt7/Oat/Otc/Pah/Pck2/Ppat/Prodh/Prodh2/Qdpr/Renbp/Sardh/Sds/Slc16a3/Slc25a44/Sult2a8/Tat/Tdo2
#> GO:0031589 Acvrl1/Agt/Anxa2/Apoa1/Apod/Bcam/Bcan/Cd63/Cib1/Clasp1/Col16a1/Col1a1/Col3a1/Coro1a/Dab2/Dbn1/Dmtn/Dock1/Egflam/Enpp2/Epdr1/Ephb1/Fbln1/Fbln2/Fermt1/Fermt3/Flna/Fn1/Fndc3b/Frem1/Iqgap1/Itga2b/Itga8/Itgad/Itgal/Itgam/Itgav/Itgb2/Itgb3/Itgb5/Itgb6/Kif14/L1cam/Lamb1/Lamb2/Lamc1/Ldb1/Lgals1/Limch1/Lims2/Lpxn/Map4k4/Marcks/Meltf/Mertk/Mkln1/Myadm/Nid2/Notch1/Npnt/Ntn4/Olfm4/Parvg/Postn/Prex1/Ptn/Ptprk/Rac2/Rcc2/Rreb1/Sirpa/Slk/Smad3/Spock2/Src/Srcin1/St6gal1/Taok2/Tek/Thbs1/Vcam1/Vwf/Zyx
#> GO:0170041 Abat/Ahcy/Aldh18a1/Aldh1a1/Aldh6a1/Arg1/Arg2/Asl/Ass1/Comt/Cps1/Cth/Dao/Dpep1/Ero1a/Gamt/Gnmt/Icmt/Ido2/Kmo/Kyat1/Kyat3/Mthfr/Otc/P4ha1/Pemt/Phgdh/Plod2/Prdx4/Sardh/Slc1a3/Srr/Tdo2/Upb1
#> GO:0071466 Abcc1/Ahr/Aim2/Akr1c12/Aox1/Cers1/Cyp1a2/Cyp1b1/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp2f2/Cyp2g1/Cyp2j11/Cyp2s1/Cyp3a11/Cyp3a41b/Cyp3a44/Cyp46a1/Dpep1/Fmo5/Gsta2/Gsta3/Gstm1/Gstm2/Gstm3/Gstm4/Gstp1/Gstp3/Kcnq2/Mcm7/Nos1/Pcna/Prkaa2/Rap2a/Recql5/Sult1b1/Sult2a1/Sult2a5/Sult2a8/Tfrc/Ugt2b1/Ugt2b37/Vkorc1
#> GO:0120254 Adh1/Adh4/Afp/Ahr/Akr1c18/Akr1c6/Aldh1a1/Aldh1a3/Alox12/Alox12b/Alox12e/Bco1/Bco2/Cyp1a2/Cyp1b1/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp2f2/Cyp2g1/Cyp2j11/Cyp2s1/Cyp46a1/Cyp4a12a/Cyp4f14/Dab2/Dgkq/Dhrs7/Fads1/Gpx1/Gstp1/Gstp3/Hsd11b2/Hsd3b4/Hsd3b5/Ppargc1a/Rbp1/Rdh9/Sdr16c5/Sdr9c7
#> GO:0072330 Abcd2/Abhd2/Acacb/Acadl/Acly/Acsbg1/Acsm2/Acsm5/Acss1/Acss2/Agt/Agxt/Akr1c18/Akr1c6/Aldh1a1/Aldh1a3/Alox12/Alox12b/Alox12e/Anxa1/Apoa4/Apoc3/Baat/Ces1f/Chst14/Cyp27a1/Cyp39a1/Cyp7b1/Degs1/Elovl2/Fabp5/Fads1/Fads2/Fasn/Gamt/Gatm/Gstm4/Gstp1/Gstp3/Hacd2/Kmo/Ldha/Ldhb/Lpgat1/Lpl/Ltc4s/Myo5a/Pdk4/Pkm/Prkaa1/Prkaa2/Prmt3/Prox1/Prxl2b/Qki/Rbp1/Rdh9/Scd1/Sds/Srr
#> GO:0006805 Ahr/Akr1c12/Aox1/Cyp1a2/Cyp1b1/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp2f2/Cyp2g1/Cyp2j11/Cyp2s1/Cyp3a11/Cyp3a41b/Cyp3a44/Cyp46a1/Fmo5/Gsta2/Gsta3/Gstm1/Gstm2/Gstm4/Gstp1/Gstp3/Nos1/Sult1b1/Sult2a1/Sult2a5/Sult2a8/Ugt2b1/Ugt2b37/Vkorc1
#> GO:0006690 Ahr/Akr1c18/Akr1c6/Alox12/Alox12b/Alox12e/Alox5ap/Anxa1/Atp6v1b1/Ces2b/Ces2c/Ces2g/Comt/Cyp1b1/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp2f2/Cyp2g1/Cyp2j11/Cyp2s1/Cyp4a12a/Cyp4f13/Cyp4f14/Dpep1/Fabp5/Fads1/Gpx1/Gstp1/Gstp3/Hpgd/Ltc4s/Ncf1/Prxl2b/Ptgr1
#> Count
#> GO:0009410 97
#> GO:0044282 97
#> GO:0046394 90
#> GO:0016053 90
#> GO:0006631 109
#> GO:1901605 66
#> GO:0006575 64
#> GO:0006790 84
#> GO:0006520 74
#> GO:0170033 52
#> GO:0007015 101
#> GO:0016054 64
#> GO:0046395 64
#> GO:0031589 83
#> GO:0170041 34
#> GO:0071466 55
#> GO:0120254 51
#> GO:0072330 60
#> GO:0006805 43
#> GO:0006690 46
head(kegg_res, 20)
#> category
#> mmu03030 Genetic Information Processing
#> mmu05204 Human Diseases
#> mmu00980 Metabolism
#> mmu00830 Metabolism
#> mmu04820 <NA>
#> mmu00982 Metabolism
#> mmu00140 Metabolism
#> mmu00620 Metabolism
#> mmu03430 Genetic Information Processing
#> mmu00480 Metabolism
#> mmu04974 Organismal Systems
#> mmu00220 Metabolism
#> mmu01230 Metabolism
#> mmu04512 Environmental Information Processing
#> mmu00983 Metabolism
#> mmu01240 Metabolism
#> mmu00380 Metabolism
#> mmu00330 Metabolism
#> mmu00052 Metabolism
#> mmu05418 Human Diseases
#> subcategory ID
#> mmu03030 Replication and repair mmu03030
#> mmu05204 Cancer: overview mmu05204
#> mmu00980 Xenobiotics biodegradation and metabolism mmu00980
#> mmu00830 Metabolism of cofactors and vitamins mmu00830
#> mmu04820 <NA> mmu04820
#> mmu00982 Xenobiotics biodegradation and metabolism mmu00982
#> mmu00140 Lipid metabolism mmu00140
#> mmu00620 Carbohydrate metabolism mmu00620
#> mmu03430 Replication and repair mmu03430
#> mmu00480 Metabolism of other amino acids mmu00480
#> mmu04974 Digestive system mmu04974
#> mmu00220 Amino acid metabolism mmu00220
#> mmu01230 Global and overview maps mmu01230
#> mmu04512 Signaling molecules and interaction mmu04512
#> mmu00983 Xenobiotics biodegradation and metabolism mmu00983
#> mmu01240 Global and overview maps mmu01240
#> mmu00380 Amino acid metabolism mmu00380
#> mmu00330 Amino acid metabolism mmu00330
#> mmu00052 Carbohydrate metabolism mmu00052
#> mmu05418 Cardiovascular disease mmu05418
#> Description
#> mmu03030 DNA replication - Mus musculus (house mouse)
#> mmu05204 Chemical carcinogenesis - DNA adducts - Mus musculus (house mouse)
#> mmu00980 Metabolism of xenobiotics by cytochrome P450 - Mus musculus (house mouse)
#> mmu00830 Retinol metabolism - Mus musculus (house mouse)
#> mmu04820 Cytoskeleton in muscle cells - Mus musculus (house mouse)
#> mmu00982 Drug metabolism - cytochrome P450 - Mus musculus (house mouse)
#> mmu00140 Steroid hormone biosynthesis - Mus musculus (house mouse)
#> mmu00620 Pyruvate metabolism - Mus musculus (house mouse)
#> mmu03430 Mismatch repair - Mus musculus (house mouse)
#> mmu00480 Glutathione metabolism - Mus musculus (house mouse)
#> mmu04974 Protein digestion and absorption - Mus musculus (house mouse)
#> mmu00220 Arginine biosynthesis - Mus musculus (house mouse)
#> mmu01230 Biosynthesis of amino acids - Mus musculus (house mouse)
#> mmu04512 ECM-receptor interaction - Mus musculus (house mouse)
#> mmu00983 Drug metabolism - other enzymes - Mus musculus (house mouse)
#> mmu01240 Biosynthesis of cofactors - Mus musculus (house mouse)
#> mmu00380 Tryptophan metabolism - Mus musculus (house mouse)
#> mmu00330 Arginine and proline metabolism - Mus musculus (house mouse)
#> mmu00052 Galactose metabolism - Mus musculus (house mouse)
#> mmu05418 Fluid shear stress and atherosclerosis - Mus musculus (house mouse)
#> GeneRatio BgRatio RichFactor FoldEnrichment zScore pvalue
#> mmu03030 24/1292 36/10571 0.6666667 5.454592 9.989876 2.940064e-14
#> mmu05204 37/1292 90/10571 0.4111111 3.363665 8.402800 3.897706e-12
#> mmu00980 33/1292 76/10571 0.4342105 3.552662 8.333493 9.424495e-12
#> mmu00830 38/1292 102/10571 0.3725490 3.048155 7.755832 6.864827e-11
#> mmu04820 63/1292 233/10571 0.2703863 2.212270 6.981959 4.556316e-10
#> mmu00982 28/1292 72/10571 0.3888889 3.181846 6.931615 7.239657e-09
#> mmu00140 32/1292 99/10571 0.3232323 2.644651 6.134719 1.122112e-07
#> mmu00620 19/1292 44/10571 0.4318182 3.533088 6.282645 2.733909e-07
#> mmu03430 13/1292 23/10571 0.5652174 4.624546 6.493078 4.481884e-07
#> mmu00480 25/1292 73/10571 0.3424658 2.802017 5.764811 8.034345e-07
#> mmu04974 32/1292 108/10571 0.2962963 2.424263 5.551268 1.022391e-06
#> mmu00220 12/1292 21/10571 0.5714286 4.675365 6.290744 1.076431e-06
#> mmu01230 26/1292 79/10571 0.3291139 2.692773 5.635097 1.163854e-06
#> mmu04512 28/1292 89/10571 0.3146067 2.574077 5.564385 1.278278e-06
#> mmu00983 29/1292 96/10571 0.3020833 2.471612 5.404682 2.106978e-06
#> mmu01240 40/1292 154/10571 0.2597403 2.125166 5.248361 2.117713e-06
#> mmu00380 19/1292 52/10571 0.3653846 2.989536 5.366417 5.644824e-06
#> mmu00330 19/1292 54/10571 0.3518519 2.878813 5.164789 1.067085e-05
#> mmu00052 14/1292 33/10571 0.4242424 3.471104 5.305021 1.324229e-05
#> mmu05418 37/1292 149/10571 0.2483221 2.031744 4.732687 1.531910e-05
#> p.adjust qvalue
#> mmu03030 1.005502e-11 6.932361e-12
#> mmu05204 6.665077e-10 4.595190e-10
#> mmu00980 1.074392e-09 7.407322e-10
#> mmu00830 5.869427e-09 4.046635e-09
#> mmu04820 3.116520e-08 2.148662e-08
#> mmu00982 4.126605e-07 2.845058e-07
#> mmu00140 5.482318e-06 3.779745e-06
#> mmu00620 1.168746e-05 8.057836e-06
#> mmu03430 1.703116e-05 1.174201e-05
#> mmu00480 2.747746e-05 1.894414e-05
#> mmu04974 3.061832e-05 2.110958e-05
#> mmu00220 3.061832e-05 2.110958e-05
#> mmu01230 3.061832e-05 2.110958e-05
#> mmu04512 3.122651e-05 2.152889e-05
#> mmu00983 4.526611e-05 3.120840e-05
#> mmu01240 4.526611e-05 3.120840e-05
#> mmu00380 1.135606e-04 7.829353e-05
#> mmu00330 2.027462e-04 1.397819e-04
#> mmu00052 2.383613e-04 1.643365e-04
#> mmu05418 2.619565e-04 1.806041e-04
#> geneID
#> mmu03030 Fen1/Lig1/Mcm2/Mcm3/Mcm4/Mcm5/Mcm6/Mcm7/Pcna/Pola2/Pold1/Pold3/Pole/Prim1/Rfc1/Rfc2/Rfc3/Rfc4/Rfc5/Rnaseh2a/Rnaseh2b/Rnaseh2c/Rpa1/Rpa2
#> mmu05204 Cyp1a2/Cyp1b1/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp3a11/Cyp3a25/Cyp3a41b/Cyp3a44/Gsta2/Gsta3/Gstk1/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Mgst1/Nat2/Sult2a1/Sult2a5/Sult2a8/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37
#> mmu00980 Adh1/Adh4/Akr1c21/Aldh3a1/Aldh3b1/Cbr2/Cyp1a2/Cyp1b1/Cyp2f2/Cyp2s1/Dhdh/Gsta2/Gsta3/Gstk1/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Mgst1/Sult2a1/Sult2a5/Sult2a8/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37
#> mmu00830 Adh1/Adh4/Aldh1a1/Aldh1a3/Aldh1a7/Aox1/Aox2/Aox3/Bco1/Cyp1a2/Cyp2a22/Cyp2a4/Cyp2a5/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2s1/Cyp3a11/Cyp3a25/Cyp3a41b/Cyp3a44/Cyp4a12a/Rdh9/Sdr16c5/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37
#> mmu04820 Ampd3/Atp1b4/Ckm/Col1a1/Col1a2/Col3a1/Col4a3/Col5a1/Col6a4/Csrp1/Csrp2/Csrp3/Des/Diaph3/Eln/Fbln1/Fbln2/Fbn1/Fhl2/Fhl3/Flnc/Fn1/Itga2b/Itga8/Itgav/Itgb3/Itgb5/Itgb6/Lbr/Ldb3/Lmnb1/Mybpc1/Mybpc3/Mybph/Mybphl/Myh10/Myh3/Myh8/Myom1/Nebl/Nid2/Pdlim2/Pdlim4/Pdlim7/Sdc1/Sgca/Sgce/Snta1/Sptbn4/Sspn/Sun1/Sun2/Thbs1/Tmod2/Tnnc2/Tnni1/Tnnt1/Tnnt3/Tpm4/Trim54/Vcan/Vim/Zyx
#> mmu00982 Adh1/Adh4/Aldh3a1/Aldh3b1/Aox1/Aox2/Aox3/Cyp1a2/Fmo5/Gsta2/Gsta3/Gstk1/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Mgst1/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37
#> mmu00140 Akr1c18/Akr1c21/Comt/Cyp1a2/Cyp1b1/Cyp2b23/Cyp2b9/Cyp2c29/Cyp2c40/Cyp2c50/Cyp2c54/Cyp2c55/Cyp2c67/Cyp2c69/Cyp2c70/Cyp2d9/Cyp3a11/Cyp3a25/Cyp3a41b/Cyp3a44/Cyp7b1/Hsd11b2/Hsd3b1/Hsd3b4/Hsd3b5/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37
#> mmu00620 Acacb/Acat1/Acat3/Acss1/Acss2/Adh1/Adh4/Aldh2/Aldh7a1/Glo1/Ldha/Ldhb/Ldhd/Mdh1/Me1/Me2/Me3/Pck2/Pkm
#> mmu03430 Lig1/Msh2/Msh6/Pcna/Pold1/Pold3/Rfc1/Rfc2/Rfc3/Rfc4/Rfc5/Rpa1/Rpa2
#> mmu00480 Ggt1/Gpx1/Gpx7/Gpx8/Gsta2/Gsta3/Gstk1/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Idh1/Lap3/Mgst1/Nat8/Nat8f2/Oplah/Prdx6/Rrm2/Sms/Txndc12
#> mmu04974 Atp1b4/Cela3b/Col12a1/Col14a1/Col16a1/Col18a1/Col1a1/Col1a2/Col2a1/Col3a1/Col4a3/Col5a1/Col6a4/Cpa1/Cpb1/Ctrb1/Ctrl/Dpp4/Eln/Mme/Prcp/Prss1/Slc16a10/Slc36a4/Slc3a1/Slc3a2/Slc7a7/Slc7a8/Slc8a2/Slc8a3/Slc9a3/Try5
#> mmu00220 Arg1/Arg2/Asl/Ass1/Cps1/Gls2/Glul/Gpt/Gpt2/Nags/Nos1/Otc
#> mmu01230 Aldh18a1/Arg1/Arg2/Asl/Asns/Ass1/Bcat1/Bcat2/Cps1/Cs/Cth/Glul/Gpt/Gpt2/Idh1/Nags/Otc/Pah/Pfkl/Pgam2/Phgdh/Pkm/Psat1/Psph/Pycr2/Sds
#> mmu04512 Cd44/Col1a1/Col1a2/Col2a1/Col4a3/Col6a4/Fn1/Frem1/Frem2/Gp1ba/Gp1bb/Gp9/Hmmr/Itga2b/Itga8/Itgav/Itgb3/Itgb5/Itgb6/Lamb1/Lamb2/Lamc1/Npnt/Reln/Sdc1/Thbs1/Tnc/Vwf
#> mmu00983 Cda/Ces1f/Ces2b/Ces2c/Ces2g/Gsta2/Gsta3/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Mgst1/Nat2/Nme7/Rrm2/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37/Upb1/Upp2/Xdh
#> mmu01240 Ak1/Ak3/Ak6/Ak7/Alas1/Aldh2/Alpl/Bcat1/Bcat2/Bco1/Coq4/Coq7/Ctps1/Ctps2/Fech/Gch1/Ggh/Haao/Hpd/Ido2/Kmo/Mthfd1l/Nme7/Nqo1/Pank3/Pdxp/Phospho2/Pnpo/Psat1/Sdr16c5/Tdo2/Ugdh/Ugt1a2/Ugt1a5/Ugt1a9/Ugt2a3/Ugt2b1/Ugt2b36/Ugt2b37/Vkorc1
#> mmu00380 Acat1/Acat3/Acmsd/Aldh2/Aldh7a1/Aox1/Aox2/Aox3/Cyp1a2/Cyp1b1/Ddc/Gcdh/Haao/Ido2/Inmt/Kmo/Kyat1/Kyat3/Tdo2
#> mmu00330 Aldh18a1/Aldh2/Aldh7a1/Arg1/Arg2/Ckb/Ckm/Ckmt1/Dao/Gamt/Gatm/Lap3/Nos1/Oat/P4ha1/Prodh/Prodh2/Pycr2/Sms
#> mmu00052 Akr1b10/Akr1b7/Akr1b8/Gale/Galk1/Ganc/Gck/Hk1/Hk2/Hk3/Hkdc1/Lct/Pfkl/Sis
#> mmu05418 Akt3/Arhgef2/Ass1/Chuk/Cyba/Gpc1/Gsta2/Gsta3/Gstm1/Gstm2/Gstm3/Gstm4/Gstm6/Gstp1/Gstp3/Gstt3/Hmox1/Itga2b/Itgav/Itgb3/Keap1/Mapk12/Mgst1/Mmp2/Ncf1/Ncf2/Nqo1/Pik3cd/Prkaa1/Prkaa2/Rac2/Sdc1/Sqstm1/Src/Sumo2/Thbd/Vcam1
#> Count
#> mmu03030 24
#> mmu05204 37
#> mmu00980 33
#> mmu00830 38
#> mmu04820 63
#> mmu00982 28
#> mmu00140 32
#> mmu00620 19
#> mmu03430 13
#> mmu00480 25
#> mmu04974 32
#> mmu00220 12
#> mmu01230 26
#> mmu04512 28
#> mmu00983 29
#> mmu01240 40
#> mmu00380 19
#> mmu00330 19
#> mmu00052 14
#> mmu05418 37